News Story
Ghodssi, Bentley patent compounds that give new way to kill drug-resistant bacteria
COLLEGE PARK, Md. – In the current era of antibiotic-resistant bacteria, fighting bacterial infections presents a difficult challenge for microbiologists and clinicians. The challenge is heightened when harmful bacteria “communicate” to form biofilms—protective matrixes of polysaccharides and proteins that encase bacteria attached to surfaces. Even when antibiotics are effective against single cells, they are oftentimes unable to eradicate the biofilm itself.
To attack this problem, University of Maryland researchers have developed chemical compounds that inhibit the formation and persistence of biofilms and enhance the effectiveness of conventional antibiotics. On Feb. 10, 2015, the researchers were awarded U.S. Patent 8,952,192 for the compounds.
The patented compounds inhibit the bacterial communication process called quorum sensing. Through quorum sensing, neighboring bacterial cells produce and detect chemical signals called autoinducers. As the number of cells in a population increases, so does the concentration of these autoinducers. When a specific concentration of autoinducers is reached, bacteria begin to act less like single cells and more like a colony. They can coordinate their gene expression and perform complex pathogenic and symbiotic processes, such as forming biofilms, producing virulence factors and inducing bioluminescence.
"We’re using bacterial cells’ own machinery against them, by blocking signals that would normally result in biofilm formation. These molecules also trick bacteria into exhibiting behaviors that will alert the immune system to their presence,” says Herman Sintim, associate professor in the Department of Chemistry and Biochemistry.
UMD co-inventors of these compounds with Sintim, include William Bentley, Fischell Department of Bioengineering Chair; Reza Ghodssi, Herbert Rabin Distinguished Chair in Engineering from theDepartment of Electrical and Computer Engineering and director of the Institute for Systems Research and a part of UMD's Brain and Behavior Initiative; Jacqueline Smith, who received her doctorate in chemistry from UMD in 2011; and Varnika Roy and Mariana Meyer, who received doctorates in bioengineering from UMD in 2011 and 2014, respectively. The compounds were created with support from the National Science Foundation (NSF), the Camille & Henry Dreyfus Foundation and UMD startup funds.
The researchers’ compounds are based on autoinducer AI-2 and its precursor DPD (4, 5-dihydroxy-2, 3-pentanedione), which is produced or recognized by more than 70 species of bacteria. The team developed analogs of DPD that do not trigger bacterial virulence, but are similar enough in structure to be recognized by DPD and AI-2 receptors. By binding with the receptors, the analogs block AI-2 from binding, preventing the receptors from sending quorum sensing signals.
In a 2010 paper published in the Journal of the American Chemical Society and a 2012 paper published in the journal ACS Chemical Biology, the researchers found that the shape and flexibility of the analog compounds are important for generating productive interactions between the analog and AI-2 receptor. Moreover, some analogs work better against certain bacterial species than others. For example, one analog called propyl-DPD (DPD plus C3H7) worked against E. coli, but only butyl-DPD (DPD plus C4H9) had an effect on Salmonella typhimurium.
“We found that disruption of AI-2 signaling in environmental containing multiple species of bacteria will likely require multiple AI-2 analogs used in combination,” said Sintim.
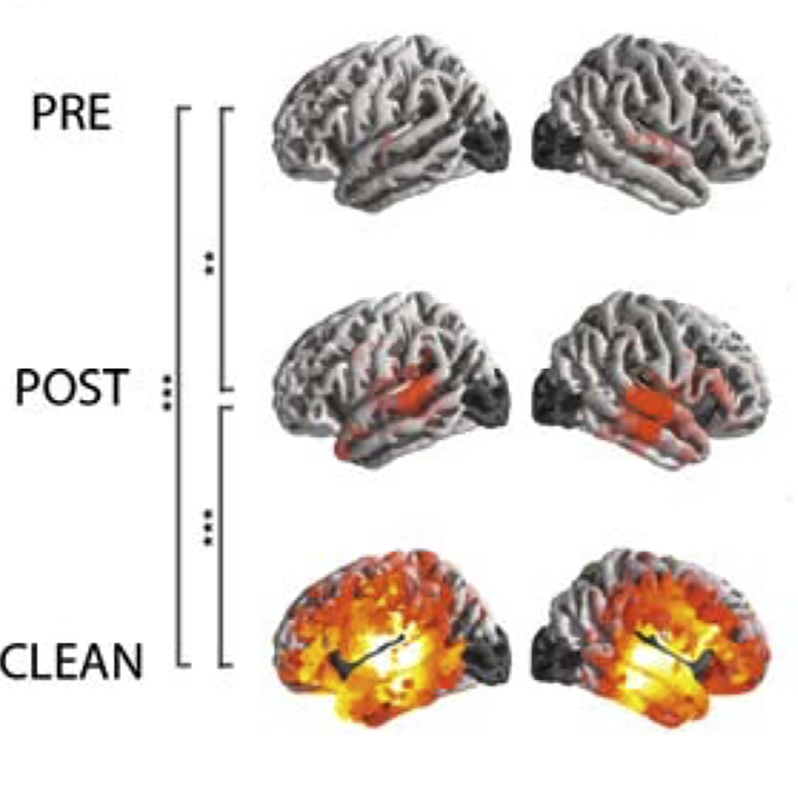
In 2013, the team demonstrated that isobutyl-DPD (DPD plus (CH3)2CH–CH2) could significantly inhibit the maturation of E. coli biofilms. Additionally, they found that combining isobutyl-DPD with the antibiotic gentamicin effectively cleared pre-existing E. coli biofilms. Similarly, phenyl-DPD (DPD plus C6H5) used in combination with gentamicin cleared pre-existing Pseudomonas aeruginosa biofilms. These results were published in the journal Applied Microbiology and Biotechnology.
Later that year, the team collaborated with Korean scientists to solve the crystal structure of a bacterial regulatory protein in complex with their synthetic analogs. Insights gained from this structural work, which was published in the Journal of the American Chemical Society, led to a new generation of stable DPD analogs that were described in the journal Chemical Communications.
“This work suggests the possibility of resurrecting traditional antibiotics, which were once effective but have been rendered ineffective due to bacterial resistance, by co-administration with innocuous AI-2-based quorum sensing inhibitors,” said Sintim.
The team’s fundamental chemical and structural findings have paved the way for them to develop more potent molecules that “tame” bacteria. The researchers are currently developing hybrid molecules combining molecules that stop bacterial communication with molecules that kill bacteria.
“Bacterial biofilms are notoriously resistant to antibiotics so putting anti-biofilm and antibacterial molecules in one hybrid unit could deliver spectacular results,” said Sintim.
Smith is now a postdoctoral researcher at Georgetown University, Meyer is a multifunction engineer in the professional development program at Northrop Grumman Corporation, and Roy is a scientist at MedImmune.
Published February 10, 2015