Molecular Connectomics of Activity-Dependent Circadian Circuit Development
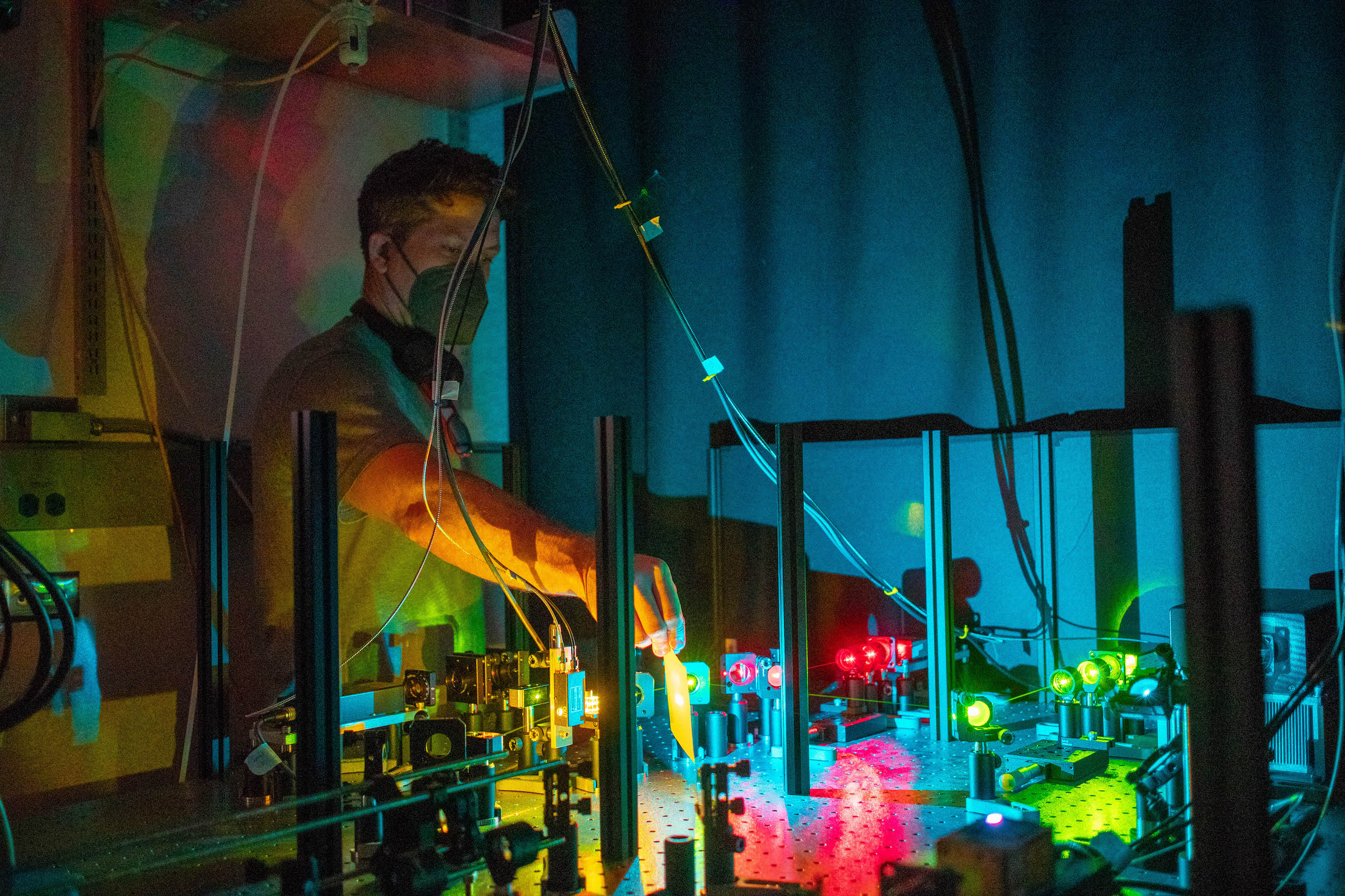
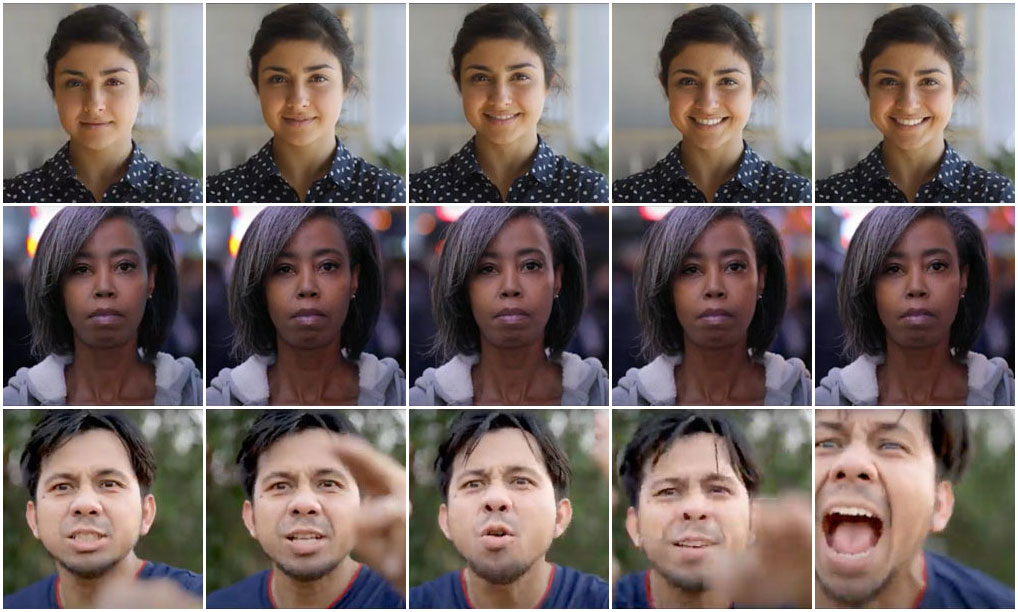
A key challenge in neurobiology is understanding how synaptic organization establishes circuit function underlying cognition and behavior. This project deploys a combination of advanced transcriptomic, proteomic, and high-resolution imaging methodologies to investigate circuit plasticity across multiple biological scales, from RNA and proteins to neurons and whole circuits.
Drs. Colenso Speer and Peter Nemes describe their BBI-funded collaboration with Dr. Najib El-Sayed.